Coronavirus Variants with Two Specific Mutations May Evade Antibodies, Says Virologist Shahid Jameel

views
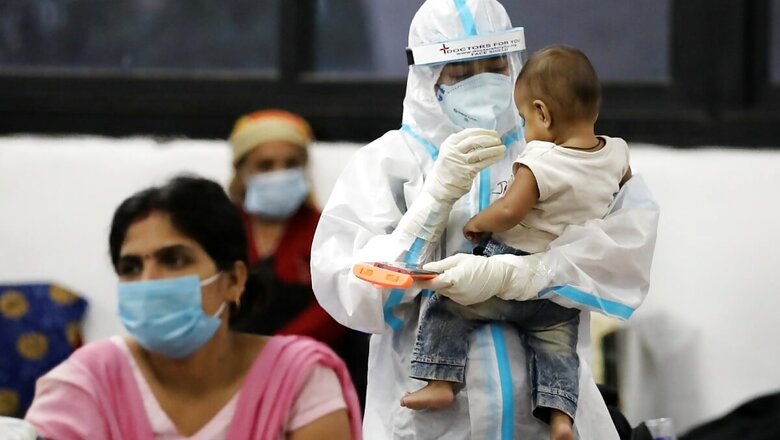
Variants of the novel coronavirus carrying two specific mutations in their spike protein may evade antibodies and make vaccines less effective, according to eminent virologist Shahid Jameel, who says the need of the hour is to improve surveillance for "homegrown" lineages of the virus. Jameel, Director of the Trivedi School of Biosciences at Ashoka University in Haryana, said these two mutations would lead to a "drastic change" in the structure of the part of the spike protein that binds with antibodies, making vaccines less effective against such variants.
The spike protein of the coronavirus enables it to bind to the ACE2 receptor on host cells — a process which studies have show to be crucial for the virus to enter the cells and cause infection. According to the noted virologist, currently, the data is consistent with poor activity of neutralising antibodies against the South African lineage of the coronavirus, which carries both these mutations on the spike protein.
At GYANTEEKA, an online presentation-discussion series hosted by the Indian National Young Academy of Science (INYAS), New Delhi to spread awareness and bust myths on vaccines, Jameel said these mutations are in the 501st and 484th amino acid building blocks that make up the virus spike protein. In his presentation on Saturday, the noted virologist said both these mutations happen at the interface where antibodies bind to the virus spike protein.
"At the interface, there are a couple of key amino acid residues. One is the N501Y mutation. Another emerging mutation is the E484K mutation, and if you notice, the 484 changes negatively charged amino acid into positively charged amino acid," he explained in his presentation, citing studies. "If you have both 501 and 484 mutations happening at the same time in the same virus, that virus is likely to either evade antibody response, or it is going to make the antibodies less effective. That's the take-home," he added.
On Tuesday, the Health ministry said four people in the country tested positive for infection with the South African variant, and one for the Brazil variant — a first for India. While in India, there have not been reports of community transmission of these virus, Jameel believes the need of the hour is also to look out for homegrown variants. "Remember India already has about 11 million infections now, so there could easily be homegrown variants and I think for that it is very important to have genomic surveillance, which is done at scale," the virologist said.
"And for that government of India has set up an inter-ministerial Group which is called INSA COG which brings together 10 different National Laboratories to start sequencing at a density of about 5 per cent," he added. Currently, the virologist said India is sequencing at a density of about 0.05 per cent.
"So essentially, the plan is to upscale sequencing about a hundredfold in the country," Jameel added. A recent study by the scientists of CSIR-Centre for Cellular and Molecular Biology (CCMB) in Hyderabad analysed over 5,000 coronavirus variants, describing how the virus has evolved over the course of the pandemic.
It found that a variant carrying the N440K mutation is spreading a lot more in the southern states of India. "We now have emerging evidence that the N440K variant is spreading a lot more in southern states. Closer surveillance is needed to understand its spread properly," CCMB Director Rakesh Mishra said in a statement.
While variants with immune-escape such as the South African variant have been identified with only a low prevalence in India, Mishra believes this could be because not enough sequencing has been done. "More coronavirus genomes need to be sequenced across the country to accurately identify the emergence of these and other new variants," he added.
.
Read all the Latest News, Breaking News and Coronavirus News here

















Comments
0 comment